by Frank Bastian
Python is a high-level, interpreted, object-oriented programming language that is often used for scientific computing due to its simple syntax and its rich ecosystem of libraries like numpy, scipy, pandas, tensorflow, … which reduces the programming effort heavily. Therefore, Python is a often-used language among members of EvoGamesPlus project.
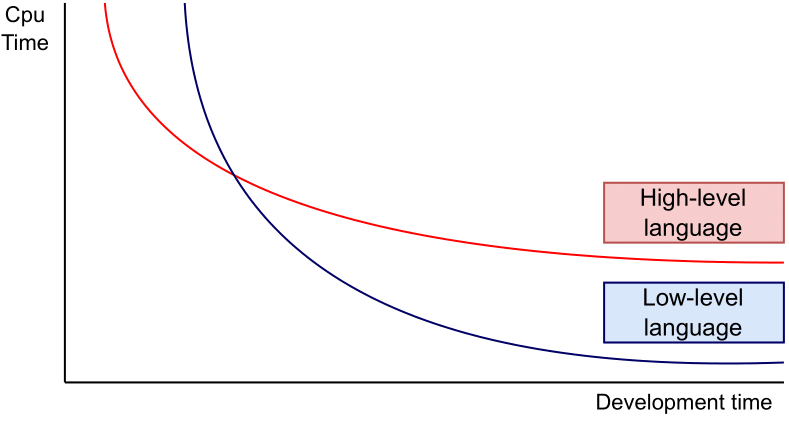
However, Python might lack computational speed compared to lower-level languages like C, C++, etc. However, using these languages is often associated with a higher programming effort. This balancing act is illustrated in Figure 1. For Python, we will demonstrate how to navigate this balancing act, either by using a just-in-time compiler or C extensions.
Compiler vs Interpreter
Conceptual programming languages can be separated into two categories: compiled and interpreted languages. The key difference between these two is “what” and “where” something is executed.
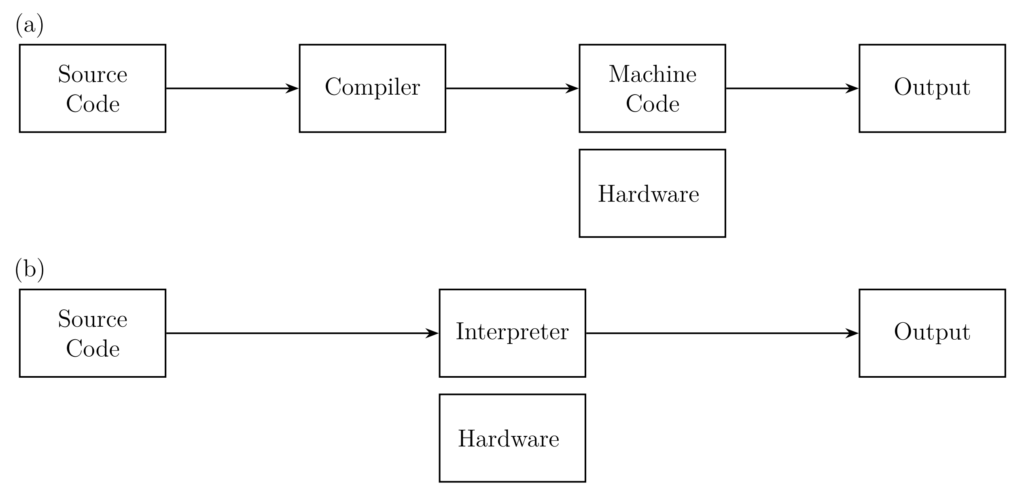
In compiled languages such as C, C++, … the source code is translated by a program called compiler into machine code (binaries), which are executed directly on hardware. The main benefit is the fast execution time of binary files through a direct execution on hardware, combined with the optimisations performed during compilation. One drawback is the lack of portability, as binary files are compiled for a particular processor architecture and operating system (OS). Hence, compiled binaries may not run “everywhere”.
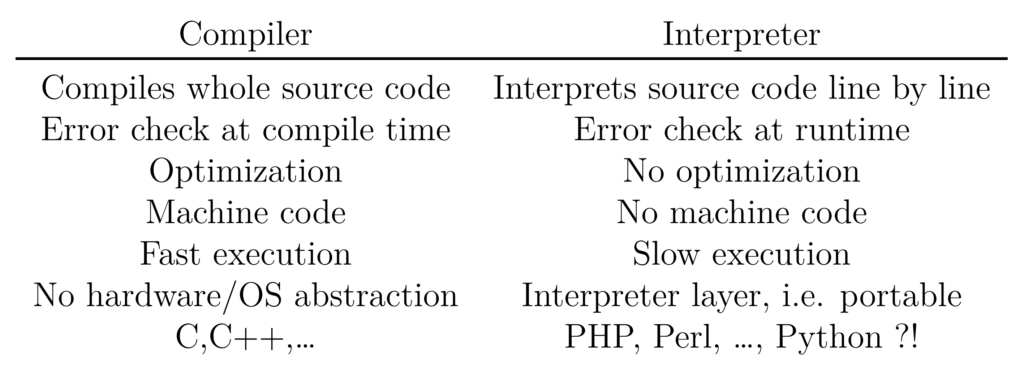
Interpreted languages address this limitation by introducing an additional abstraction layer called an interpreter. An interpreter is a program that runs on the local OS and interprets (i.e. executes) the given source code. Using this additional abstraction layer, the program has become independent of hardware architectures and operating systems can be run nearly “everywhere”. However, interpreted languages have the disadvantage of being slower than their compiled counterparts due to the lack of compiler optimisation and the extra layer. A short comparison of both concepts is given in Table 1 while a scheme of interpreter and compilers can be found in Figure 2. Modern programming languages, especially the interpreted ones, are starting to combine the advantages of both concepts by using just-in-time compilation, i.e. compiling the source code immediately before it is executed.
Next, we illustrate the usage of such just-in-time compiler.

Numba
Numba is a just-in-time compiler for Python. It attempts to compile annotated Python functions that have been decorated using @jit to low level code. If Numba successfully compiles the annotated functions, an increase in speed up to one or two orders of magnitude can be achieved according to the documentation [2]. Furthermore, using Numba often requires only minimal changes to the existing code base, and in contrast to the approach in the next section, no further knowledge of a language like C is required.

Since we only want to raise awareness of the existence of Numba we refer for more details to their documentation.
C extension
Alternatively to the usage of a just-in-time compiler like Numba, it is possible to create a C extension for Python as a shared library.
C File
First, the computationally expensive part needs to be rewritten in C. Important is the keyword extern for C linkage. Furthermore, for Windows, the functions need to be explicitly exported to a dynamic-link library (dll). In our example, this is achieved via a preprocessor macro checking for the target operating system.

While it is possible to manually compile the C source code using gcc or MSVC, it is much more convenient (and less error prone) to use the library setuptools via a Python script. Setuptools we will call the corresponding compiler with all necessary flags and create the desired shared library (.so Linux and .pyd Windows).
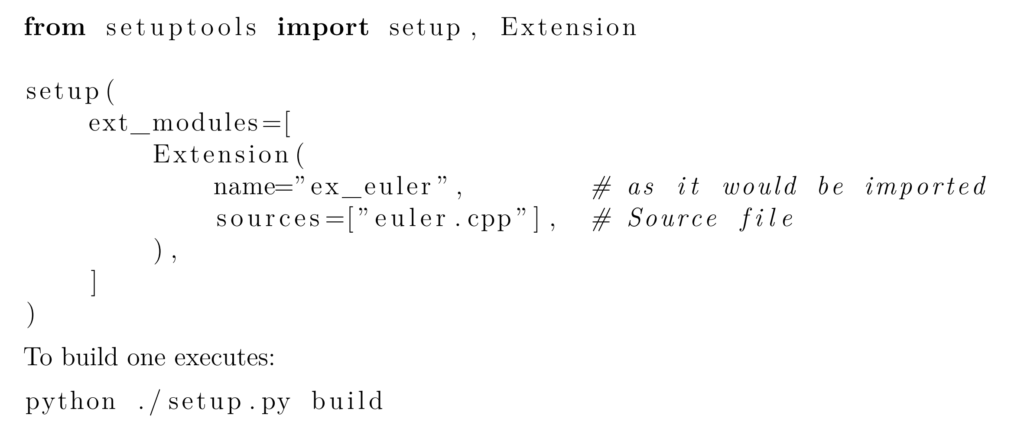

Remark: For Windows the dll (the .pyd file) has to contain an initialization function PyInit_foo() that will be called by using import foo [3,4].
C Wrapper
To use the previously created shared object via Python, we rely on the library ctypes. First, ctypes needs to load the shared object. Afterwards, the signature of the implemented C-method, i.e. arguments and return types, has to be specified within the attributes argtypes and restype. A comprehensive list of all corresponding C data types can be found in the documentary [5]. It is also possible to pass an array, respectively the pointer to the array. However, it is the programmer’s responsibility to prevent indexing out of bounds by defining the expected shape.

Conclusion
We conclude by comparing run time performance and implementation effort of the three presented implementations that are
- Vanilla Python,
- Numba,
- C extension.
As illustrated in Figure 1, the development effort of the C extension is significantly higher than plain Python code of small adjustments such that it can be compiled by Numba. For the benchmark we assume h=0.0001 and T=150, i.e. 1500000 Euler steps. Vanilla Python needs 9.9 seconds, Numba 0.59 seconds, including compilation, and the C extension 0.12 seconds. Clearly, the C extension outperforms Python and Numba in execution time. Therefore, if execution time is important it can be highly beneficial to write the computationally expensive tasks as a C extension.
References
[2] url: https://numba.readthedocs.io/en/stable/.
[3] url: https://stackoverflow.com/questions/34689210/error-exporting-symbol-when-building-python-c-extension-in-windows.
[4] url: https://docs.python.org/3/faq/windows.html.
[5] url: https://docs.python.org/3/library/ctypes.html.